 ITru_R1_5: 5'- AATGATACGGCGACCACCGAGATCTACAC NNNNNNNN ACACTCTTTCCCTACACGACGCTCTTCCGATCT -3' Either way, they are generally structured as follows: The second set of primers, commonly referred to as indexing primers, are normally purchased in a kit or designed in-house. The second PCR uses the Read 1 and Read 2 primer sequences as templates to add the P5 and P7 sequencing primers, as well as the i5 and i7 indexes. Note that in some sequencing protocols such as those used for whole-genome, metagenomic, or metatranscriptomics, the Read 1 and Read 2 adapters are annealed to the molecules using alternatives to PCR such as tagmentation or ligation. TruSeq Read 1 Forward primer Reverse primer TruSeq Read 2 Tailed R primer: 5'- GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT- GTRATWGCHCCDGCTARWACWGG -3'įollowing amplification with these tailed primers, the molecules will look like this:ĥ'- ACACTCTTTCCCTACACGACGCTCTTCCGATCT GGDACWGGWTGAACWGTWTAYCCHCC-Target- CCWGTWYTAGCHGGDGCWATYAC AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC -3'ĥ'- TGTGAGAAAGGGATGTGCTGCGAGAAGGCTAGA CCHCCYATWTGWCAAGTWGGWCADGG-Target- GGWCAWRATCGDCCHCGWTARTG TCTAGCCTTCTCGTGTGCAGACTTGAGGTCAGTG -3' Tailed F primer: 5'- ACACTCTTTCCCTACACGACGCTCTTCCGATCT- GGDACWGGWTGAACWGTWTAYCCHCC -3' In the below example we are using the fwhF2-fwhR2n primer sets which amplify a short region of the mitochondrial COI barcode: To achieve this we need to modify our locus-specific primers to include the Universal 5' adapters as tails. The first PCR amplifies the target DNA and adds the illumina Read 1 primer on the left side of the insert, and the Read 2 primer on right side of insert. In our current metabarcoding protocol, we are using the TruSeq adapter system and anneal them to the molecule using 2 separate PCRs: First PCR: Illumina now encourages customers to use unique dual indexing (UDI) whenever possible to ensure the most accurate demultiplexing, and therefore reduce the risk of sample cross-contamination. Or completely unique (Unique Dual Indexing) where both the i5 and i7 index is completely unique to that sample: Dual indexed libraries can either be combinatiorial, where only 1 index is different between samples, while the other is shared: Most modern sequencing protocols use dual-indexing rather than single indexing. The index at the right hand side is the "i7 index", or "index1", and the index at the left hand side is the "i5 index", or "index2". CTATGTTA) that are unique to each sample. The "N"s in the above diagrams indicate the "indexes", or "barcodes" used to discriminate different samples. The "insert" is the commonly used term for the target DNA that is to be sequenced, in the case of metabarcoding libraries this also includes the forward and reverse PCR primers used to amplify the target DNA. Illumina P5 i5 Next era Read 1 Next era Read 2 i7 Illumina P7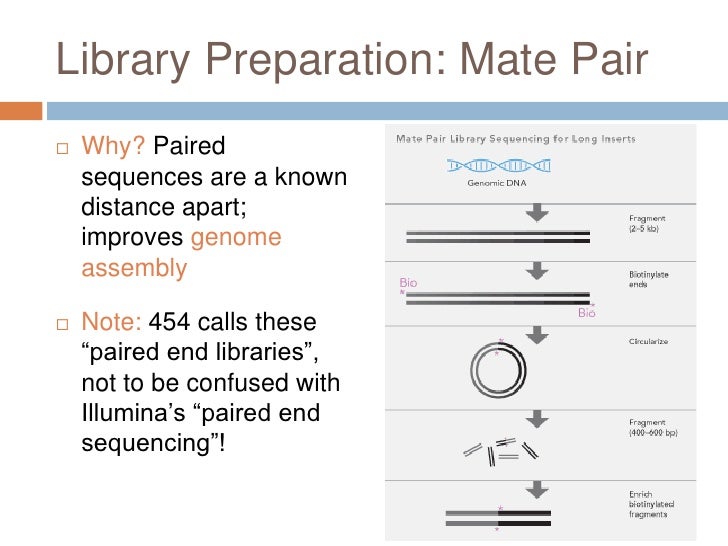 
Illumina P5 i5 TruSeq Read 1 TruSeq Read 2 i7 Illumina P7 Nextera Dual Index Library:ĥ'- AATGATACGGCGACCACCGAGATCTACAC NNNNNNNN TCGTCGGCAGCGTC AGATGTGTATAAGAGACAG-insert- CTGTCTCTTATACACATCT CCGAGCCCACGAGAC NNNNNNNN ATCTCGTATGCCGTCTTCTGCTTG -3'ģ'- TTACTATGCCGCTGGTGGCTCTAGATGTG NNNNNNNN AGCAGCCGTCGCAG TCTACACATATTCTCTGTC-insert- GACAGAGAATATGTGTAGA GGCTCGGGTGCTCTG NNNNNNNN TAGAGCATACGGCAGAAGACGAAC -5' There are a variety of Illumina and third party adapter designs that can be used for Illumina sequencing, with the TruSeq and Nextera adapter systems being the most popular: TruSeq Dual Index Library:ĥ'- AATGATACGGCGACCACCGAGATCTACAC NNNNNNNN ACACTCTTTCCCTACACGACGCTCTTCCGATCT-insert- AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC NNNNNNNN ATCTCGTATGCCGTCTTCTGCTTG -3'ģ'- TTACTATGCCGCTGGTGGCTCTAGATGTG NNNNNNNN TGTGAGAAAGGGATGTGCTGCGAGAAGGCTAGA-insert- TCTAGCCTTCTCGTGTGCAGACTTGAGGTCAGTG NNNNNNNN TAGAGCATACGGCAGAAGACGAAC -5' (2) The i5 and i7 index sequences (barcodes) which uniquely label the molecules from different samples to allow multiplexing/pooling of multiple samples in a single sequencing run or flow cell lane.(3) The binding sites for the Read 1 and Read 2 sequencing primers which initiate the sequencing process itself. 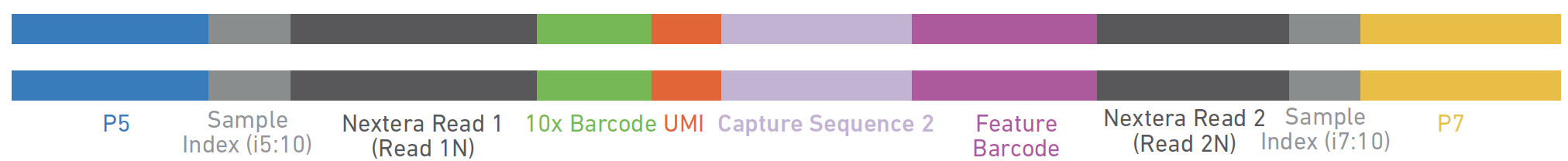 
These adapters consist of three main components: (1) the P5 and P7 sequences that allow the library to bind and generate clusters on the flow cell. Illumina sequencing by synthesis requires special oligonucleotide adapters to be annealed to the purified target DNA in order to initiate sequencing. Illumina sequencing Illumina sequencing libraries 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
